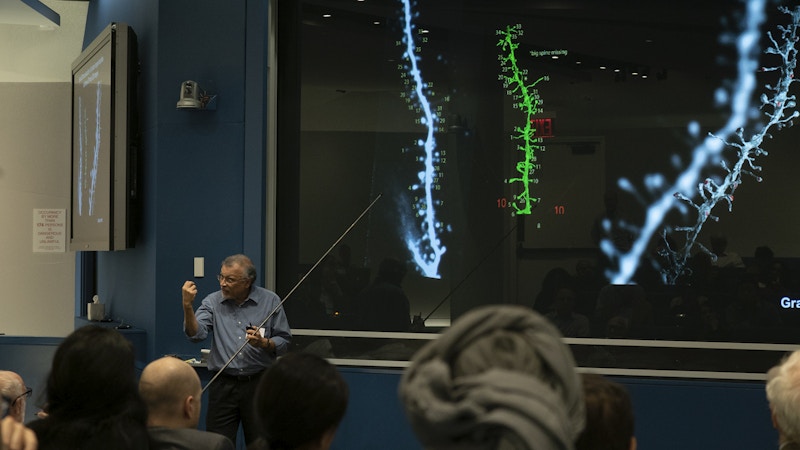
The Simons Foundation Autism Research Initiative (SFARI) held its fourteenth science meeting September 30–October 2, 2018. Over 100 SFARI Investigators, collaborators and foundation staff gathered to discuss recent findings and future directions of SFARI-funded studies and other research relevant to autism spectrum disorder (ASD).
Keynote presentations
The meeting opened with a keynote address by Randy Buckner, who presented his research on the functional organization of the cerebral cortex in human and nonhuman primates1,2. Discussing the relevance of these studies in the context of autism research, he emphasized the need for experiments that bridge the gap between known genetic changes in ASD and their effects on the development and function of cortical association networks that regulate social cognition, language and other higher-order processes affected in the disorder. He emphasized that studies at the single-subject level may complement group-average investigations by providing insights into the anatomical variability of the large-scale association networks. He also highlighted the importance of studying higher-order processes, or their proto-form, in animal models. One way to move the field forward, in Buckner’s opinion, is to make association circuits tractable so they can be targeted in intervention studies.
The Monday night keynote lecture was presented by Dinakar Singh and Loren Eng, who are the founders of the Spinal Muscular Atrophy (SMA) Foundation as well as parents of a woman with this disorder. A rare condition associated with mutations in SMN1 and SMN2 genes, SMA leads to progressive loss of neuromuscular control and is a major genetic cause of death in children. Singh and Eng recounted their emotional journey since their daughter was first diagnosed and discussed the challenges of establishing the SMA Foundation at a time when little was known about this condition and no treatment was available. Today, their efforts are starting to pay off, as the foundation is a leading funder of SMA research and the first FDA-approved drug for SMA, SPINRAZA® (nusinersen), has proven successful in increasing the likelihood of overall survival and improving motor function3,4. With several other treatments under development, the couple plans to continue their battle to find novel interventions to give their daughter, and others similarly affected, the chance for a better life.
Session presentations
Seventeen SFARI Investigators presented their latest research findings during the four sessions on the topics of genetics; molecular mechanisms; circuits, systems, cognition & behavior; and clinical studies.
Genetics
The genetics session focused on key questions related to the genetic basis of ASD, including the relationship between genotypic and phenotypic variability, and the role that different genetic factors, such as de novo and inherited mutations, play in autism risk.
Genotype to phenotype
The first speaker, Graeme Davis, hypothesized that phenotypic penetrance in ASD may be determined by the complex interaction between genetic changes and mechanisms that regulate neuronal function, such as homeostatic synaptic plasticity. In particular, he argued that presynaptic homeostatic plasticity may provide a ‘physiological buffer’ to the disorder, which, when disrupted, would increase the system’s susceptibility to genetic and environmental insults, and contribute to ASD phenotypic severity. A goal of Davis’ research is thus to elucidate the links between homeostatic plasticity and ASD genetics. In his presentation, he described data from the model species Drosophila that suggest that interactions between ASD risk genes and the rest of the genome may be associated with dysfunction of presynaptic homeostatic mechanisms.
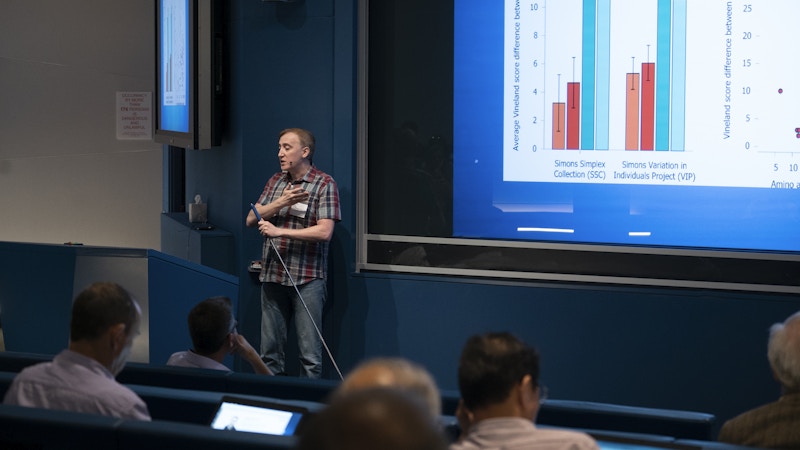
The next presentation also focused on genetic-phenotypic relationships in ASD and related neurodevelopmental disorders. Dennis Vitkup discussed a recent study from his lab that examined the impact of likely gene-disrupting (LGD) de novo mutations on phenotypes, such as intelligence quotients, in unrelated probands of the Simons Simplex Collection (SSC) and Simons Variation in Individuals Project (Simons VIP). Findings from this study suggest that while probands with LGD mutations in the same gene showed more similar phenotypes relative to other probands in these cohorts, the greatest phenotypic similarities were found between individuals harboring LGD mutations in the same exon5. These results may indicate that exons constitute a ‘unit of effective phenotypic impact,’ which is key to shaping phenotypic variation in ASD and related neurodevelopmental disorders.
Common and rare variants
Daniel Geschwind’s presentation centered around the genetic architecture of ASD. He discussed results from a collaboration with Dennis Wall that is part of the Hartwell Autism Research and Technology Initiative (iHART). Whole-genome sequencing data from more than 400 hundred multiplex families from the Autism Genetic Resource Exchange (AGRE) were analyzed, resulting in the identification of inherited and de novo mutations in 16 new genes associated with ASD risk6. Geschwind also described findings from ongoing analyses that are assessing the role of noncoding portions of the genome in ASD risk. He concluded his talk with a description of a recent functional genomic study that examined development of the human cerebral cortex at the single-cell level7.
The session ended with a presentation by Elise Robinson who provided an overview of the last decade of autism genetics research and discussed future directions for the field. In particular, she argued that inherited polygenic risk is likely to be an important contributing factor to ASD in addition to rare variation, as shown by a recently published study led by Anders Børglum and Mark Daly8 (Figure 1). Robinson also discussed the NeuroDev study9, an initiative she is leading in collaboration with the Broad Institute, the Kenyan Medical Research Institute, Oxford University and the University of Cape Town. This project aims to collect whole blood for exome sequencing from children with neurodevelopmental disorders and their parents in Africa, as well as associated cognitive behavioral information, with the goal to increase diversity in autism cohorts.

Molecular mechanisms
The next session, which was on molecular mechanisms, delved deeper into the biochemical processes that underlie neuronal dysfunction in ASD. The presentations also included discussions of recent advances in methodologies, including the generation of stem cells and brain organoids, that are being used to model biological changes associated with the disorder in vitro.
Discussing his research on the mechanisms regulating postsynaptic densities, the first speaker, Mingjie Zhang, described a platform that his lab recently developed to study synaptic structure and plasticity10. Applications of this tool to autism research may help elucidate the effects of genetic mutations on synaptic function, which is known to be affected in ASD. In particular, Zhang’s research aims to understand how mutations in SHANK3, a high-confidence autism risk gene, affect interactions of key proteins in postsynaptic densities.
Continuing the discussion on biochemical mechanisms that regulate neuronal signaling, Kristopher Kahle described findings on a novel kinase-regulated phosphorylation switch of NKCC1/KCC2 transporters. He showed that this switch underlies the developmental process whereby GABA changes from modulating excitatory to inhibitory signaling in the brain. He also demonstrated that experimentally tuning off this mechanism restores GABAergic inhibition to hyperexcitable circuits in a mouse model of Rett syndrome.
The last two speakers in this session, Kevin Eggan and Lilia Iakoucheva discussed their research using induced pluripotent stem cells (iPSCs) and brain organoids to better understand functional mechanisms that are altered in ASD. With a focus on methodological aspects, Eggan described new techniques he developed in collaboration with Steven McCarroll that allow iPSCs from hundreds of individuals to be examined at the same time, thus, reducing the costs and experimental noise typically associated with producing and analyzing individual cell lines. In addition, he discussed experiments he is conducting in collaboration with SFARI that aim to better understand variability among iPSC lines generated from distinct source cells with different reprogramming methods.
Iakoucheva then presented data from a study assessing the functional impact of the 16p11.2 deletion/duplication copy number variant (CNV), one of the most frequent among the CNVs conferring ASD risk, on brain development. By creating brain organoids from fibroblasts of 16p11.2 deletion and duplication carriers collected from the Simons VIP, she found that these in vitro systems recapitulated aspects of brain growth observed in individuals with 16p11.2 duplication/deletion. In addition, she identified a number of molecular and cellular signatures associated with the syndrome.
Circuits, systems, cognition & behavior
The session on circuits, systems, cognition & behavior included studies using animal models as well as human research studies to investigate circuits that may be altered in ASD.
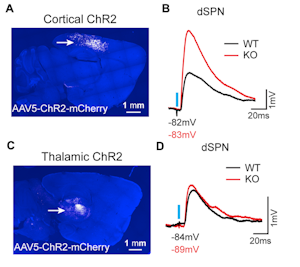
The first speaker in this session, Helen Bateup, presented her lab’s recent work on TSC1, a syndromic autism risk gene that plays a role in regulating mTOR complex 1 (mTORC1). A viral strategy was used to postnatally delete Tsc1 from a subset of mouse striatal neurons. Loss of Tsc1 resulted in an increase in the excitability of striatal spiny projection neurons of the direct pathway (dSPNs) but not the indirect pathway (iSPNs). In addition, a selective enhancement of corticostriatal transmission onto dSPNs lacking Tsc1 was found (Figure 2), highlighting the critical role of mTORC1 in regulating striatal activity in a cell-type and input-specific manner11. Using a similar approach, her lab also found that activation of mTORC1 due to the loss of Tsc1 from mouse midbrain dopamine neurons disrupted dopamine release and the structure of axonal terminals, and led to deficits in cognitive flexibility12. Bateup’s studies aim to elucidate how genetic changes lead to alterations in neural circuits that may underlie repetitive and inflexible behaviors in ASD.
Mriganka Sur then discussed his research on Arc, an immediate early gene that is upregulated in excitatory neurons by synaptic activity. He described findings from a recent study examining synaptic plasticity in the primary visual cortex of mice and showed that Arc regulated synaptic plasticity by modulating the local interplay of potentiated and depressed spines13. As Arc is a key regulator of a number of signaling pathways implicated in ASD, understanding its role in shaping neuronal plasticity may help to shed light on mechanisms underlying the disorder.
In addition to repetitive and inflexible behaviors and synaptic deficits, abnormal processing of sensory stimuli has also been frequently associated with ASD. Stelios Smirnakis presented preliminary findings in a mouse model of MECP2 duplication syndrome — a condition characterized by intellectual disability, autism, spasticity, severe epilepsy and early death — which suggests alterations in visual processing. In particular, he highlighted differences between wildtype and mutant mice in responding to visual stimuli at different contrast levels and found that responses in the primary visual cortex of the mutants were more reliable than in wildtype mice. Smirnakis argued that his findings may suggest alterations in brain canonical functions that have been previously predicted to be affected in
ASD (e.g.,14).
Dora Angelaki further discussed this theoretical framework14 and hypothesized that one form of canonical computation, causal inference, would be affected in ASD. She presented preliminary data from studies that aim to test this hypothesis by asking participants with and without ASD to perform tasks on multisensory integration that either require or do not require causal inference. Angelaki’s computations are also consistent with the hypothesis of an increased ratio of neural excitation to inhibition (E/I) in ASD15 — a finding that she discussed in greater detail at a SFARI workshop focused on the E/I hypothesis.
Finally, the session concluded with Ethan Scott, who described his recent work using whole-brain imaging of neuronal activity in the visual, auditory16 and vestibular17,18 systems of zebrafish. Earlier in the year, Scott attended a SFARI workshop where he and others discussed the advantages and challenges of using zebrafish as experimental models to study neurodevelopmental disorders. Building on his studies of wildtype zebrafish, he now aims to investigate the neurobiology of circuits underlying multisensory integration, and their dysfunction, in mutant zebrafish harboring ASD risk variants.
Clinical studies
The last session of the meeting was focused on clinical studies. Presentations in this session described human research studies investigating the neural bases of ASD and assessing ASD phenotypes, as well as preliminary results from therapeutic intervention studies.
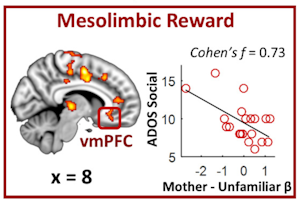
Opening this session, Vinod Menon presented findings from an imaging study aimed at understanding the basis of lack of social interest typically seen in individuals with ASD. He showed that structural and functional connectivity of key structures of the mesolimbic reward system is altered in children with ASD relative to typically developing children and that those changes correlated with parent-reported measures of social interaction impairments19. Menon went on to show that children with autism also showed alterations in brain activity of the mesolimbic reward areas and other regions while listening to unfamiliar voices as well as their mother’s voice, one of the most rewarding and instrumental stimuli for social-emotional development during childhood20 (Figure 3). He concluded that his research lends support to a ‘social motivation theory’ of autism, which postulates that individuals with autism may interact less socially because they find social interactions less rewarding than neurotypical individuals21, 22.
The next speaker, Pawan Sinha, discussed studies that he is conducting to test another hypothesis of ASD, the so-called ‘predictive impairment hypothesis.’ Originally proposed by Sinha and colleagues several years ago23, this hypothesis argues that ASD is associated with impairments in cognitive processes that require estimation of conditional probabilities (i.e., understanding B given A). At the meeting, he provided evidence from studies on sensory habituation, as well as studies on motor functions he is conducting in collaboration with Dagmar Sternad that support an account of the disorder based on a ‘predictive impairment.’ Sinha and Sternad recently gave a Simons Foundation public lecture on this topic (Video 1).
In addition to discussing key processes affected in ASD, this session also considered novel developments in pharmacological interventions. While no drug treating the core symptoms of the disorder is currently available, several research efforts have been recently directed to investigating the effects of a GABAB agonist, arbaclofen, which has shown some effects in pre-clinical studies. Timothy Roberts gave a preview of the ongoing analyses of a magnetoencephalography (MEG) study aimed at examining the effects of acute administration of arbaclofen on M100, an electrophysiological signature of auditory processing that he previously demonstrated to be altered in children with ASD24. The arbaclofen used in this study was provided by Clinical Research Associates, L.L.C. (CRA), an affiliate of the Simons Foundation. CRA has also provided the drug for a number of other animal and human studies, including two clinical trials in Europe and Canada. Roberts’ project aims to test whether arbaclofen rescues the aberrant M100 signature in individuals with ASD and to identify suitable candidates for clinical trials on the basis of an objective physiological measure.
The session ended with a presentation by Ilan Dinstein, who described the ongoing research he is conducting at the Negev Autism Center in Israel. The center brings together scientists from the university and clinicians from the nearby hospital and sees children with autism for whom a number of behavioral, speech, eye-tracking, neuroimaging and other types of data, as well as biospecimens for genetic analysis (e.g., saliva, blood) are collected. Dinstein’s research focuses on sleep, which is known to be affected in a significant proportion of individuals with ASD. He argued that sleep disturbances may be associated with sensory sensitivities observed in children with ASD25. In his talk, he presented data from overnight electroencephalography recordings comparing sleep patterns and architecture in typically developing and ASD children. Weaker slow-wave activity was observed during the first half of the night in the ASD group, suggesting that ASD children exhibit less pressure to sleep compared to typically developing children. This reduction in slow-wave activity correlated with the severity of sleep problems in the participating children, possibly indicating a specific mechanism for sleep disturbances in some cases of ASD. One goal of the center, and of Dinstein’s research, is to develop objective measures of behaviors using technologies such as eye tracking and motion tracking to help relate neurophysiological changes with core aspects of ASD phenotypes and provide reliable ways to assess outcomes after treatment interventions.
Demo sessions
In addition to the keynote and speaker sessions, the meeting also featured a number of demo sessions that provided an opportunity for researchers to learn more about informatics tools for sharing, visualizing and analyzing data sets relevant for autism research.
These sessions included the presentation of an updated version of SFARI Base, the portal to request access to SFARI data and biospecimens, as well as presentations of two online platforms that allow users to browse, visualize and analyze SFARI genomics data. These platforms are the Genotypes and Phenotypes in Families (GPF) and SFARI Viewer, which were developed, respectively, by Ivan Iossifov and Frameshift, in collaboration with the SFARI informatics team.
In addition to these SFARI resources, a Simons Foundation genomics tool, HumanBase, was also showcased. This interactive platform, which was developed by Olga Troyanskaya’s group at the Flatiron Institute (a division of the Simons Foundation), applies machine learning algorithms to massive data sets, including genomic, gene expression and protein-protein interaction data, to help researchers identify novel, data-driven associations.
“This was an intellectually stimulating meeting,” says Louis Reichardt, SFARI director. “I deeply enjoyed meeting with the investigators who attended and learning about the exciting progress in their research. Witnessing the passion and commitment of these scientists to autism research really gives us hope that we are one step closer to better understanding the disorder and on the right path to developing interventions that could improve people’s lives.”
For additional information about SFARI-funded investigators and research projects, please visit: https://www.sfari.org/research/investigators.
References
- Braga R.M. and Buckner R.L. Neuron 95, 457-471 (2017) PubMed
- Buckner R.L. and Margulies D.S. Nat. Commun. 10, 1976 (2019) PubMed
- Finkel R.S. et al. Engl. J. Med. 377, 1723-1732 (2017) PubMed
- Scoto M. et al. Gene Ther. 24, 514-519 (2017) PubMed
- Chiang A.H. et al. bioRxiv (2018) Preprint
- Ruzzo E.K. et al. bioRxiv (2018) Preprint
- Polioudakis D. et al. bioRxiv (2018) Preprint
- Grove J. et al. Nat. Genet. 51, 431-444 (2019) PubMed
- De Menil V. et al. Neuron 101, 15-19 (2019) PubMed
- Zeng M. et al. Cell 174, 1172-1187 (2018) PubMed
- Benthall K.N. et al. Cell Rep. 23, 3197-3208 (2018) PubMed
- Kosillo P. et al. bioRxiv (2018) Preprint
- El-Boustani S. et al. Science 360, 1349-1354 (2018) PubMed
- Rosenberg A. et al. Proc. Natl. Acad. Sci. USA 112, 9158-9165 (2015) PubMed
- Rubenstein J.L. and Merzenich M.M. Genes Brain Behav. 2, 255-267 (2003) PubMed
- Vanwalleghem G. et al. J. Comp. Neurol. 525, 3031-3043 (2017) PubMed
- Favre-Bulle I.A. et al. Nat. Commun. 8, 630 (2017) PubMed
- Favre-Bulle I.A. et al. Curr. Biol. 28, 3711-3722 (2018) PubMed
- Supekar K. et al. Brain 141, 2795-2805 (2018) PubMed
- Abrams D.A. et al. Elife 8, e39906 (2019) PubMed
- Dawson G. et al. Child Dev. 73, 700-717 (2002) PubMed
- Chevallier C. et al. Trends Cogn. Sci. 16, 231-239 (2012) PubMed
- Sinha P. et al. Proc. Natl. Acad. Sci. USA 111, 15220-15222 (2014) PubMed
- Gandal M.J. et al. Biol. Psychiatry 68, 1100-1106 (2010) PubMed
- Tzichinsky O. et al. Mol. Autism 9, 22 (2018) PubMed





















