This fall, the recipients of the Simons Foundation Autism Research Initiative (SFARI) Bridge to Independence (BTI) Award gathered with SFARI staff and advisors for a virtual ‘retreat’ of six weekly webinars. Beginning on October 1, 2020, each of the webinars included scientific presentations by the fellows, followed by breakout sessions to discuss the practical skills needed to become successful Principal Investigators in today’s scientific ecosystem.
Launched in 2015, the BTI Award program provides autism research funding to early-career scientists who are transitioning from a mentored (e.g., postdoctoral fellowship) to an independent research position at a North American academic institution. The program currently includes 31 fellows, many of whom have already secured a faculty position and opened their independent laboratory as Principal Investigators.
“The pandemic has hit young investigators particularly hard, “noted SFARI senior scientist and BTI program lead Alice Luo Clayton as she welcomed the group. “We decided to run a virtual version of the retreat hoping that it will help the fellows continue to build the BTI community and network with senior investigators on their research and careers during these challenging times.”
New avenues in autism genetics
Despite advances in autism gene discovery, only a fraction of autism cases have an identifiable genetic cause. Most of these genetic discoveries have been based on de novo mutations found through exome sequencing; however, other types of mutations are likely to increase autism risk, such as recessive mutations. Ryan Doan of Boston Children’s Hospital showed that recessive loss-of-function and missense mutations are highly present in autism1; he is now expanding his search for risk factors to non-coding portions of the genome that control gene expression, including promoters2 and enhancers. To this end, he has developed a panel designed to examine mutations across thousands of neuron-specific promoters, which has revealed novel risk factors in whole genome and exome sequences of over 4,000 families with autism.
Variants in non-coding regions could also alter transcript splicing and isoform usage, which in turn could contribute to autism risk. Michael Gandal of the University of California Los Angeles described his efforts to characterize the full spectrum of transcript isoforms in the developing brain, as disruptions to these have been found in autism and other psychiatric disorders3. Using new long read technology that allows sequencing a transcript’s full length, Gandal has discovered novel isoforms in fetal brain with cell type-specific resolution. In conjunction with transcriptome-wide gene expression profiles, he has also identified subclasses of cells based on isoform expression, identified more specific cell type markers, and identified switches in isoform expression in development. He argued that unraveling the genetic complexity that controls isoform regulation and cell specificity may help to better understand the molecular mechanisms underlying neurodevelopmental and psychiatric conditions.

Elliott Robinson of Cincinnati Children’s Hospital Medical Center is developing a gene therapy for neurofibromatosis type 1, a condition stemming from mutation to the NF1 gene, which leads to diverse consequences, including skin abnormalities, cognitive deficits, attention deficit hyperactivity disorder (ADHD) and sometimes autism. Dopamine transmission is also altered in mice lacking a copy of NF14. Because NF1 is so large — 350 kb — adeno-associated viral vectors used for gene therapy would not be able to deliver the full gene. Robinson described ways to get around this, including delivery of a smaller part of the gene to substitute and upregulating the remaining working copy of the NF1 gene via CRISPR activation.
Vulnerabilities during development
Autism risk genes are highly expressed during mid-fetal development in the prefrontal cortex (PFC) in humans — a time when cortical neurogenesis is peaking5. To understand these genes’ role in autism risk, several BTI fellows are focusing their research on aspects of early brain development.
Tomasz Nowakowski of the University of California San Francisco has been studying how cells acquire their identities during development in human brain tissue6,7. By transcriptome profiling thousands of single cells, he has found that neurons of the prefrontal and visual cortex show gene expression differences already at post-mitotic stages of development. In contrast, by measuring chromatin accessibility, he recently discovered an even earlier divergence among intermediate progenitor cells. Analysis of the active sites in chromatin also suggested that retinoic acid signaling is key for specifying PFC identity8.
Rebecca Muhle of Columbia University presented multiple approaches her lab is taking to understand the function of CHD8, a high-confidence autism risk gene encoding a chromatin remodeler that regulates the availability of DNA sequence to transcription factors. This has widespread downstream effects, with Muhle reporting that CHD8 binds to promoters of about 13,000 genes in human neural stem cells. In mouse embryos missing a copy of CHD8, Muhle used single-cell RNA sequencing to show that autism risk genes and CHD8 targets are enriched in maturing neurons of the developing cortex. Changes in gene transcription were mild but could still induce a ripple effect to impact brain function throughout life. Muhle is conducting this study in collaboration with SFARI Investigator James Noonan.
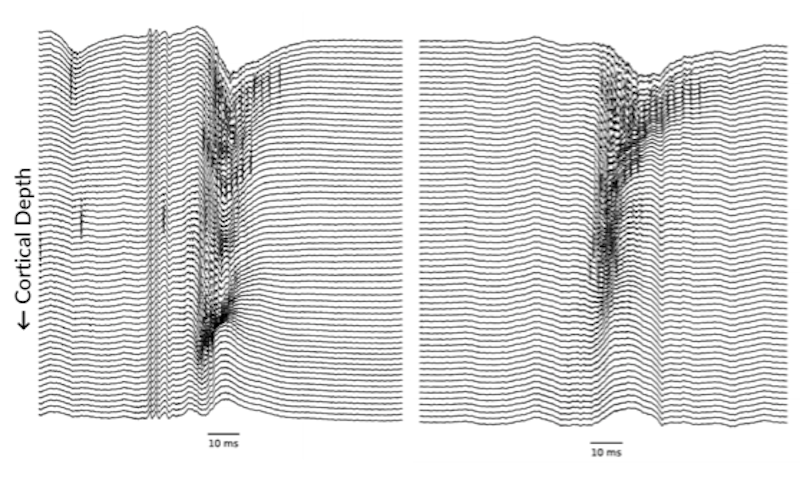
Renata Batista-Brito of Albert Einstein College of Medicine has been working to understand how information processing may be shaped by MEF2C, a transcription factor that, when disabled by mutation in humans, results in intellectual disability and features of autism. Notably, MEF2C is found specifically in parvalbumin (PV) interneurons, which are fast-spiking neurons that act as a crucial inhibitory damper on excitatory activity in the brain. Batista-Brito is exploring how embryonic removal of MEF2C in mice affects inhibitory circuitry in vivo, and how this in turn shapes visual processing and behavior. This research also tracks how state — quiet or active — modulates visual responses.
To get the single cell-level details of events that unfold in the developing fetal brain, Seth Shipman of the University of California San Francisco and the Gladstone Institutes has been developing tools that can create, within a cell, a log of its gene expression during development. His earlier work determined that the CRISPR-Cas system could be harnessed to store gene expression events, written with DNA in the order that they occur9,10. This molecular recorder relies on a reverse transcriptase to encode and store information about gene expression; Shipman is now optimizing this system to find efficient reverse transcriptases, with a goal of reading out the expression history of 5–20 genes in a single cell.
Circuits and behavior
To understand how the brain processes social information, a function that is often affected in autism, Nicholas Frost of the University of California San Francisco has recorded activity across multiple neurons simultaneously in freely behaving mice. During social interactions, neurons in the medial prefrontal cortex modulated their activity — some up, some down — leading Frost to investigate whether the temporal organization of the activity mattered. It did: using a neural network to classify patterns of activity as happening during social interactions or home cage time, Frost found that correlations between ensembles of neurons were key to transmitting social information, even when levels of activity were maintained. This type of information transmission was disrupted in SHANK3 knockout mice, a model of autism. These findings suggest that changes in the temporal dynamics of prefrontal neurons may be implicated in some of the social challenges associated with the condition11.
In addition to difficulties in social interaction, people with autism often experience challenges in many other domains, including sleep. Noting the importance of sleep to health, and the consistent reports of disrupted sleep in autism, Graham Diering of the University of North Carolina Chapel Hill has been exploring the interaction of sleep with genetic risk in mice carrying disabling mutations to SHANK3, a synaptic gene linked to autism. Diering has found that introducing sleep deprivation early in postnatal life can result in social deficits in SHANK3 mutant mice not seen otherwise. This suggests that sleep disruption can potentiate effects of gene mutation, and drive lasting behaviors. To better understand the mechanisms underlying this finding, Diering is also using proteomic analysis to explore shifts in protein production that occur with sleep deprivation in juveniles.
Insights from other species
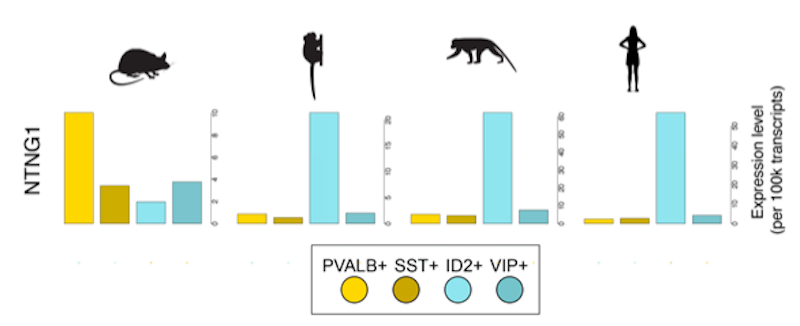
To bridge genetics to biology, autism researchers are using diverse animal models. Conducting single-cell RNA sequencing to profile gene expression in individual cells to pin down cell type identity, Fenna Krienen of Harvard Medical School has tracked changes in interneuron abundance, gene expression, allocation and type across mouse, ferret, macaque, marmoset and human brains12. She showed that interneuron number and properties differ across species — a result that not only illustrates how brains evolve, but that also highlights the importance of choosing the appropriate animal models for autism studies based on the research questions. She now plans to leverage this knowledge to study the effects of SHANK3 haploinsufficiency on cell types and circuits relevant to ASD across rodent and primate species.
Other fellows are using invertebrate model species to help understand the basic biology underlying autism. Ethan Greenblatt of the University of British Columbia made the case that Drosophila oocytes could help reveal a fundamental function of the FMR1 gene, mutations in which may lead to fragile X syndrome. In earlier work, he has found that the oocytes rely on FMR1 to maintain their competence during quiescence13. In this period no transcription occurs, but previously deposited mRNAs are translated — a scenario resembling that in neurons, where mRNA stored in neural processes is translated into protein. He found that FMR1 loss leads to oocyte failure, and a prominent decrease in translation for many of its targets, especially those with long transcripts, many of which are linked to intellectual disability in humans.
Anomalies in neuron structures like spines have been identified in autism, and these could alter how information is processed in the brain. Glia may have a hand in governing neural structure, according to work from Aakanksha Singhvi of Fred Hutchinson Cancer Research Center. She presented her research on a specific glia-neuron interaction in the nematode Caenorhabditis elegans in which a glial cell ensheathing the bushy ending of a thermosensory neuron can affect its shape, function and the animal’s sensory behavior14. Recent work suggests that the glial cell actively prunes away microvilli from the neuron, in a process that may be activity-dependent, and involve molecules already linked to autism15.
Career roundtables
After the scientific presentations, the BTI fellows gathered in small groups in virtual breakout rooms to get career advice from BTI advisors about key personal skills of successful PIs, such as leadership and time-management, as well as on key practical skills such as recruiting lab members, getting grants and managing a budget. The advisors were Ted Abel of the University of Iowa, Aravinda Chakravarti of New York University School of Medicine, Moses Chao of New York University School of Medicine, Wendy Chung of Columbia University and the Simons Foundation, Kamran Khodakhah of Albert Einstein College of Medicine, Genevieve Konopka of the University of Texas Southwestern Medical Center, David Lewis of the University of Pittsburgh, Kelsey Martin of the University of California Los Angeles, Carol Mason of Columbia University, Brian O’Roak of Oregon Health & Science University, Louis Reichardt, former SFARI director, Huntington Willard of Genome Medical, and Mark Zylka of the University of North Carolina at Chapel Hill.
Highlights included the need to stay focused on a small number of scientific projects, yet to remain flexible within their bounds; the need to hire people for their scientific personalities and willingness to learn, rather than solely on the techniques they know; exploring different styles of leadership; to not hesitate in spending startup money, which has been given to help a new investigator succeed; to safeguard time as a precious commodity; and the importance of self-care throughout this time of intense career development. Personal anecdotes from advisors provided especially useful lessons.
BTI fellow Stephanie Rudolph of Albert Einstein College of Medicine noted that she had filled a notebook with career pointers. “This is secret advice that one only hears in conversation,” she said.
This year, the unexpected challenges brought by the COVID-19 pandemic made these conversations even more compelling. Fellows noted the benefits of the BTI program in providing funding that could be used during pandemic-induced lab closures that have tied up institutional funds meant for starting a lab. The meeting was also a useful opportunity to present research results and get feedback from leaders in the field. “Under normal circumstances, I would be traveling to meetings to present our unpublished work, and receiving feedback that would help me write better papers and inspire future research directions. Due to the COVID pandemic, I am unable to do that,” Nowakowski said.
Despite the virtual format, these roundtables provided a way for fellows to connect candidly with each other and with advisors. They felt too short to some fellows, and others noted that the weekly recurrence helped with building relationships and a sense of community with each other. This could provide a source of support for the fellows, as well as potential collaborations. Future topics for the next meeting (hopefully an in-person one) could include how to manage service expectations as young faculty and how to reach out to the autism community to aid research.
The retreat was enjoyable for the advisors as well. Willard noted that the BTI retreat exposed him to new ideas percolating among young investigators and is a useful reminder of what it is like to start over amid an ever-shifting scientific landscape. “This means that there is no right way or wrong way to begin a career,” he said. He also counseled patience: “It takes time to develop a cadence and process of running a lab, and with the BTI program, they’re off to a great start.”
References
- Doan R. N. et al. Nat. Gen. 51, 1092-1098 (2019) PubMed
- An J-Y. et al. Science 362, eaat6576 (2018) PubMed
- Gandal M. et al. Science 362, eaat8127 (2018) PubMed
- Robinson J.E. et al. Elife 8, e48983 (2019) PubMed
- Willsey A.J. et al. Cell 155, 997-1007 (2013) PubMed
- Nowakowski T.J. et al. Science 358, 1318-1323 (2017) PubMed
- Cadwell C.R. et al. Neuron 103, 980-1004 (2019) PubMed
- Ziffra R.S. et al. bioRxiv (2020) Preprint
- Shipman S.L. et al. Science 353, aaf1175 (2016) PubMed
- Shipman S.L. et al. Nature 547, 345-349 (2017) PubMed
- Frost N.A. et al. bioRxiv (2020) Preprint
- Krienen F.M. et al. Nature 586, 262-269 (2020) PubMed
- Greenblatt E. et al. Science 361,709-712 (2018) PubMed
- Singhvi A. et al. Cell 165, 936-948 (2016) PubMed
- Raiders S. et al. bioRxiv (2020) Preprint


