Because of the coronavirus pandemic, the Simons Foundation Autism Research Initiative (SFARI) ran the fall 2020 science meeting virtually in a series of six webinars, beginning on October 2, 2020. Each Friday, SFARI Investigators presented their latest findings in autism research, ranging from autism genetics, through molecular mechanisms and neural circuits, to clinical insights.
Over 300 people attended the meeting, between SFARI Investigators and their lab members, and SFARI staff and collaborators.
One benefit of an online version of the meeting is that more people could attend than the in-person meeting, including graduate students and postdoctoral fellows in SFARI’s Investigators laboratories.
“It’s hard to recreate some of the spontaneous interactions and fun of the in-person meeting” said SFARI interim director John Spiro welcoming attendees. “But we are certainly going to try.”
Genetics
Mark Daly of the University of Helsinki gave an update on autism gene discovery, which has already made strides since the publication of a substantial crop of genes found through exome sequencing1. By adding new data from SPARK (Simons Foundation Powering Autism Research for Knowledge) to their samples, Daly and colleagues implicated 71 genes in autism risk with genome-wide significance. Notably, 61 of the 71 genes were already associated with developmental conditions/intellectual disability, and eight were associated with schizophrenia. The findings raise interesting questions about participant phenotypes, mutation types and shared biology — and show that integrating vast datasets across disorders will be helpful for better understanding the biological causes that are shared or unique to these conditions.
To date, much autism gene discovery has been fueled by the study of de novo mutations, the spontaneously occurring events thought to arise in zygotes. But Joseph Gleeson of the University of California San Diego has been examining the possibility that they can also come from a father’s sperm. Using deep whole-genome sequencing of sperm and blood samples, Gleeson has detected de novo mutations in some — but not all — sperm (‘mosaicism’). This finding suggests that quantifying the frequency of mutations in sperm could help estimate the risk of autism recurrence for families2. Their contribution is in fact substantial: of 11 pathogenic de novo mutations found in autism probands, three were also detected in the father’s sperm, affecting 0.5 percent of sperm in one dad to 14 percent in another.
Once genetic changes are identified in autism, it remains challenging to pinpoint their effects. This is especially true for missense mutations, which may or may not alter how a protein works. To better understand the impact of missense mutations, Haiyuan Yu of Cornell University, together with Kathryn Roeder of Carnegie Mellon University and Bernie Devlin of the University of Pittsburg, has developed computational methods to model the network of interactions between proteins (‘interactome’) and how these are disrupted by mutation3. Applying this framework to de novo missense mutations, Yu found that interactome-disrupting mutations occur more frequently in autism probands than in their unaffected siblings. These mutations tended to affect ‘hub’ genes encoding proteins that interact with many other proteins. Of 34 hub proteins identified, 12 are known autism proteins, and the others, Yu suggested, may turn out to be linked to autism as well4.
Genetic risk factors from non-coding regions of the genome were also discussed. David FitzPatrick of the University of Edinburgh described his research focused on mutations in cis-regulatory elements (CREs), such as enhancers or repressors, that might explain cases of X-linked intellectual disability. Given that the CREs on the X chromosome have been identified5, FitzPatrick and his team sequenced these for 48 families, and ultimately identified six variants — all rare, three never seen before and one that controlled FMR1, the gene linked to fragile X syndrome. Engineering the latter CRE variant into mice resulted in altered FMR1 expression in embryos and a mixed pattern of postnatal behaviors, some of which — but not all — matched phenotypes noted in FMR1 knockout mice. This suggests that CRE mutations result in misregulation of a gene, rather than a straightforward loss-of-function, making it difficult to prove a causal role for any one variant in an individual family.
Molecular mechanisms
Also focusing on FMR1, Joel Richter of University of Massachusetts Medical School described a new role for this gene. FMR1 encodes FMRP, an RNA-binding protein linked to synaptic plasticity and mRNA translation. Richter and his team found that FMRP acts as a brake for ribosomes: with an assay they developed to track ribosome movement along mRNAs, they showed that ribosomes moved quickly and generated excessive protein in hippocampal tissue in FMRP-deficient mice6. Among the RNAs specifically affected by FMRP-dependent stalling is SetD2, a protein involved in modifying chromatin. In FMRP-deficient mice, SetD2 levels increased, which in turn shifted alternative splicing, including in autism-related genes.
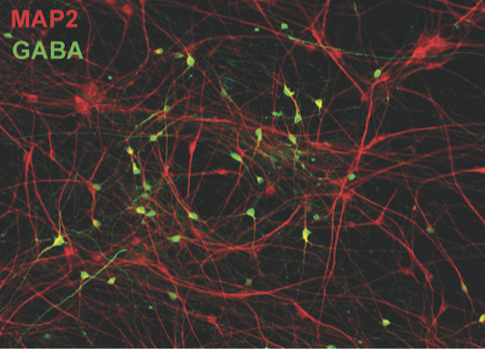
Nael Nadif Kasri of the Donders Institute presented work aimed at understanding how the action of chromatin-remodeling genes can ultimately alter neural network activity. Kasri has been studying neurons derived from people with Kleefstra syndrome (KS), which is caused by loss-of-function mutations of a chromatin-methylating gene called EHMT1 and is associated with autism. By growing excitatory neurons from induced pluripotent stem (iPS) cells on multi-electrode arrays, Kasri found enhanced NMDA receptor signals and bursting in networks from cells derived from individuals with KS syndrome compared to those from people without this condition7. Adding inhibitory neurons to these cultured networks further altered burst features. These network phenotypes can also be used to explore the effects of other genes linked to autism, such as CHD138, ANK2 or KMT2D (Figure 1).
Mechanistic insights about the relationship between chromatin remodeling and autism can come from studies of the BAF complex, a research focus of Gerald Crabtree at Stanford University. This complex regulates gene accessibility and is composed of assemblies of 15 subunits that are combined in hundreds of different ways. Mutations in genes encoding these subunits are linked to autism, including BAF250B (also known as ARID1B) and BAF53B, a neuron-specific subunit whose mutation causes Mendelian recessive autism. Crabtree has been engineering iPS cells to carry BAF250B loss-of-function mutations, growing them into neurons, then examining their effects on gene expression. This has revealed a link between BAF250B dosage and HOX gene expression, which is important in establishing cell type during brain development. With BAF250B haploinsufficiency, however, this process is undermined, potentially skewing cell type development.
Nenad Sestan of Yale University has been working to understand the mechanisms behind the specialized development of human prefrontal cortex, one brain region that is affected in autism. Gene expression profiling of human brain tissue has revealed differences in gene expression patterns between emerging brain regions during fetal development, a time and place where autism-related genes are particularly active9. Sestan noted that the genes driving these expression differences are related to retinoic acid (RA), which suggests the metabolite helps set up brain patterning and specialization. Consistent with this result, he found higher RA levels in human fetal brain than in macaque or mouse brain, as well as a lateral expansion of the RA-synthesizing enzyme, ALDH1A3, in humans10.
Circuits and behavior
Disruptions to the diverse set of ASD genes may lead to similar outcomes at the level of brain circuit function. To screen cortical networks efficiently, Jessica Cardin of Yale University and her collaborators Michael Higley and Michael Crair have developed a way to manipulate genes in cells across the brain via whole-brain adenovirus infection. She showed activity across the cortex in awake, behaving mice as they transition from a quiescent state to an aroused, ambulatory state11. Cardin used CRISPR technology to selectively knock out the Rett syndrome-linked gene MECP2 in either excitatory neurons or inhibitory neurons early in post-natal life. When the mice transition from a quiescent state to an active state, both excitatory and inhibitory neurons missing MECP2 were robustly activated, but functional connectivity between pairs of cortical regions and network structures were disrupted.
Noting that autism is marked by altered sensitivity to sensory stimuli, Bilal Haider of the Georgia Institute of Technology has been investigating visual processing and perception in mouse models of autism. Using a task in which mice indicate their detection of a visual stimulus, Haider’s team has been able to chart their contrast sensitivity while also recording neural activity from visual cortex12. In mice missing both copies of the autism risk gene CNTNAP2, Haider’s team found impairments in stimulus detection speed, accuracy and contrast sensitivity. Simultaneous recordings during the visual detection task found increased activity in inhibitory neurons and decreased activity in excitatory neurons in the mutant mice relative to the wild type. Also, Haider suggested that low frequency oscillations in layer 2/3 might degrade stimulus perception, particularly in the mutant mice13.
To understand FMR1 function, Geoffrey Goodhill of the University of Queensland has been tracking development, behavior and sensory coding in zebrafish missing FMR1, which could provide some insight into the altered sensory processing apparent in autism (Movie 1). Zebrafish are advantageous models because they show measurable behaviors just days after birth, and their entire nervous system can be non-invasively imaged. Goodhill reported that zebrafish missing both FMR1 copies show altered craniofacial development, temporary deficits in their ability to capture paramecium to eat and delayed retinotectal map development, which may indicate an abnormal reliance on peripheral vision. In an assay of social behavior, fish missing both FMR1 copies showed more enthusiasm for social interaction than wild-type fish did, though the mutant fish were slower to respond to a social partner.
Because SCN2A loss-of-function mutations have been linked to autism and intellectual disability, Kevin Bender of the University of California San Francisco (UCSF) has been exploring ways to remedy this loss in mouse models. SCN2A encodes the sodium channel Nav1.2, which Bender’s lab has found modulates dendritic excitability, synaptic function and synaptic plasticity in mice14. In collaboration with Nadav Ahituv at UCSF, Bender is now seeking ways to activate promoters for SCN2A through CRISPR technology, so as to increase expression of the remaining working copy of SCN2A and compensate for the lost allele.
CHD8 regulates transcription of many genes and has emerged as a high confidence autism risk gene. To understand how changes in transcription translate into altered brain circuitry, Laura Andreae of King’s College London has been examining the synaptic physiology of mice lacking a copy of CHD8. Recording from brain slices of the prefrontal cortex, her team has identified anomalies at both excitatory and inhibitory synapses, including altered compensation in response to a loss of inputs. The work highlights the need to track down the origins of synaptic dysfunction — whether it arises as a cell-autonomous consequence of, or network adaptation to, a gene mutation.
Clinical
Prospective studies of infant siblings of children with ASD have demonstrated that atypical brain development precedes the appearance of the defining behavioral symptoms of the condition (e.g.,15). Within-family recurrence of ASD is known to be mediated by inherited genetic variation and predicted by autism severity in the older sibling with ASD (proband)16. Using a large cohort from the Infant Brain Imaging Study (IBIS), Joseph Piven of the University of North Carolina at Chapel Hill and colleagues have found proband autism severity explained–variation in early postnatal cerebral volume, cortical surface area and white matter microstructure of the splenium among infants who were later diagnosed with ASD. These findings converge on cortical regions and fiber pathways that suggest the infant visual system may play a notable role in the downstream development of ASD and that familial indices of genetic risk for ASD may help elucidate the pathways from genetic risk, to early brain development, to disorder.
As part of the Autism Innovative Medicine Studies-2-Trials (AIMS-2-Trials), Emily Jones of Birkbeck University of London has been looking for ways to meaningfully parse the heterogeneous cases of autism using electroencephalogram (EEG) signals. She focused on the N170, an EEG signal that is slowed in response to faces in people with autism compared to people without autism17. This may well index aspects of social function, as Jones described that N170 latency correlates with brain responses to faces, genetic risk burden for autism and change in everyday social behavior, as measured by the Vineland Adaptive Behavior Scales social score. Jones suggested that N170 latency might eventually help identify appropriate participants for clinical trials.
References
- Satterstrom F.K. et al. Cell 180, 568-584 (2020) PubMed
- Breuss M.W. et al. Nat. Med. 26, 143-150 (2020) PubMed
- Chen S. et al. Nat. Genet. 50, 1032-1040 (2018) PubMed
- Chen S. et al. Mol. Autism 11, 76 (2020) PubMed
- Naville M. et al. Nat. Commun. 6, 6904 (2015) PubMed
- Shah S. et al. Cell Rep. 30, 4459-4472 (2020) PubMed
- Frega M. et al. Nat. Commun. 10, 4928 (2019) PubMed
- Mossink B. et al. bioRxiv (2020) Preprint
- Willsey A.J. et al. Cell 155, 997-1007 (2013) PubMed
- Shibata M. et al. bioRxiv (2019) Preprint
- Barson D. et al. Nat. Methods 17, 107-113 (2020) PubMed
- Speed A. et al. Cell Rep. 26, 2868-2874 (2019) PubMed
- Del Rosario J. et al. Cereb. Cortex Epub ahead of print (2021) PubMed
- Spratt P.W.E. et al. Neuron 103, 673-685 (2019) PubMed
- Hazlett H.C. et al. Nature 542, 348-351 (2017) PubMed
- Girault J.B. et al. J. Neurodev. Disord. 12, 5 (2020) PubMed
- Kang E. et al. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 3, 657-666 (2018) PubMed


