Gene discovery in autism spectrum disorder (ASD) has produced a number of promising ASD risk loci in recent years. However, these loci tend to reflect alterations within gene-coding regions, encompassing only 1.5 percent of the total genome. With the advent of new sequencing and data-analysis technologies, it is now becoming technically feasible and affordable to assess variations within the vast expanse of noncoding DNA as well.
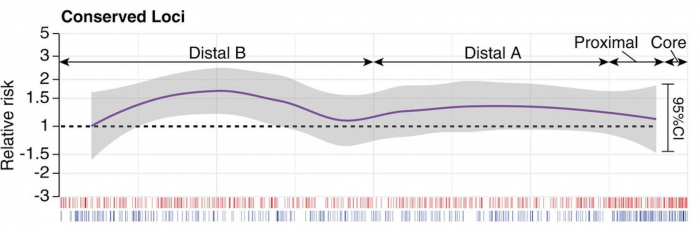
SFARI Investigator Stephan Sanders and colleagues recently found that noncoding single nucleotide variation (SNV) mutations contribute to ASD risk and provide a compelling demonstration that noncoding risk variants can be uncovered through an examination of whole-genome sequencing (WGS) data. By taking advantage of the large repository of WGS data collected as part of the Simons Simplex Collection (SSC), Sanders and colleagues examined 7,608 genomes from 1,902 family quartets for de novo SNVs present in affected children but not their unaffected siblings. The availability of this large SSC WGS data set and an analysis approach that relied on a combination of machine learning tools and comparative data analyses (i.e., a category-wide association study framework) allowed the authors to extract meaningful associations from the wealth of over 255,000 de novo mutations identified across the WGS data set, uncovering a significant effect of noncoding mutations within gene-promoter regions, particularly within evolutionarily conserved transcription factor binding sites, in children with autism.
The demonstration that de novo noncoding SNVs alter risk for ASD fits nicely with a study from SFARI Investigator Jonathan Sebat demonstrating a similar increase in parentally inherited noncoding structural variant burden in ASD based on an analysis of 2,644 ASD families from the SSC (Brandler et al., Science, 2018). Olga Troyanskaya, a Simons Foundation Flatiron Institute scientist, also found both transcriptional and posttranscriptional noncoding variation in an analysis of the SSC and other cohorts (Zhou et al., bioRxiv, 2018). Two additional studies from SFARI Investigator Evan Eichler’s laboratory using SSC and other WGS cohorts have reported similar trends toward an increase in noncoding de novo ASD mutations (Turner et al., Am. J. Hum. Genet., 2016; Turner et al., Cell, 2017). Finally, work from SFARI Investigators Ivan Iossifov and Michael Wigler suggests that there is an increased burden of small insertions and deletions in the introns of SSC probands (Munoz et al., bioRxiv, 2017).
Combined, these studies point to a significant burden of ASD risk from noncoding mutations and further highlight a need for very large cohort sizes in order to identify the specific genes affected by them.
Reference(s)
Genome-wide de novo risk score implicates promoter variation in autism spectrum disorder.
An J.Y., Lin K., Zhu L., Werling D., Dong S., Brand H., Wang H.Z., Zhao X., Schwartz G.B., Collins R.L., Currall B.B., Dastmalchi C., Dea J., Duhn C., Gilson M.C., Klei L., Liang L., Markenscoff-Papadimitriou E., Pochareddy S., Ahituv N., Buxbaum J., Coon H., Daly M., Kim Y.S., Marth G., Neale B.M., Quinlan A., Rubenstein J., Sestan N., State M., Willsey A. J., Talkowski M., Devlin B., Roeder K., Sanders S.