
Research Highlights
Highlights of SFARI-funded papers, selected by the SFARI science team.


Recent SFARI-funded research challenges the idea that autism represents a single condition driven by a uniform set of genetic risk factors. Instead, the results support a developmental framework in which distinct constellations of polygenic influences are linked to different diagnostic timelines.
Subscribe to our newsletter and receive SFARI funding announcements and news

SFARI-funded research demonstrates that activating serotonergic 5HT2c receptors leads to an increase in intracellular calcium, early-response gene expression and elevated protein secretion into cerebrospinal fluid.

SFARI-funded research finds that RAPGEF4 is a key molecular regulator orchestrating prefrontal cortical maturation in primates.
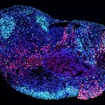
SFARI-funded research demonstrates that SCN8A mutations underlying DEE-13 disrupt both cortical and hippocampal networks, but in distinct ways.



Large-scale genetic studies have now uncovered more than 200 genes that can be linked to neurodevelopmental disorders (NDDs) or autism. Many of those genes are implicated through gene-damaging heterozygous de novo mutations that result in haploinsufficiency of the affected gene. Identifying such genes opens the door for the development of gene-specific therapeutics like gene replacement therapy. However, gene replacement therapy comes with many challenges, one of them being the risk of overexpression. Therefore, therapeutics that could result in the upregulation of the remaining healthy allele and do not interfere with endogenous regulatory mechanisms of gene expression would be ideal.

Social behavior is a complex and dynamic process shaped by movement, coordination and physical touch. A new study in Cell written by members of SFARI's Autism Rat Consortium introduces s-DANNCE, a machine-learning system that can map the fine-scale movements of freely interacting rats in three dimensions. By applying s-DANNCE to seven genetic rat models of autism, researchers have uncovered distinct social phenotypes, offering new insights into the diversity of autism-related behaviors.
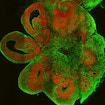
Two studies by different research groups — one led by Flora Vaccarino and the other by Juergen Knoblich — used brain organoids derived from pluripotent stem cells of people with autism and showed how transcriptional alterations affecting certain cell types during human brain development could contribute to the early emergence of ASD.
- Previous Page
- Viewing
- Next Page