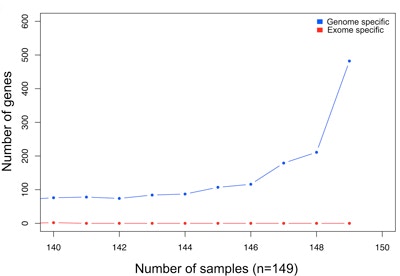
Whole-exome sequencing of families affected by simplex autism has yielded tremendous insight into genetic variants conferring risk in coding regions of the genome. Genetic variation within noncoding and regulatory regions of the genome, however, remains largely unknown. In this study, SFARI Investigator Evan Eichler and his colleagues performed whole-genome sequencing analyses of 208 genomes from 53 families from the Simons Simplex Collection (SSC). Comparison of whole-genome and whole-exome sequencing data suggests that a combined approach works best to identify single-nucleotide variants and indels, whereas the greatest benefit of whole-genome sequencing was in the discovery of small copy number variants that cannot be reliably detected through exome sequencing. The authors also reported an enrichment of variants in putative noncoding regulatory elements in individuals with autism compared with controls, specifically flanking genes that had already been implicated by de novo coding mutations. Whole-genome sequencing data from the entire SSC of 2,644 families will soon be available (see here for more details), bringing with it the promise that many more coding and noncoding variants contributing to autism risk will be discovered.
Reference(s)
Genome sequencing of autism-affected families reveals disruption of putative noncoding regulatory DNA.
Turner T., Hormozdiari F., Duyzend M.H., McClymont S.A., Hook P.W., Iossifov I., Raja A., Baker C., Hoekzema K., Stessman H., Zody M.C., Nelson B.J., Huddleston J., Sandstrom R., Smith J.D., Hanna D., Swanson J.M., Faustman E.M., Bamshad M.J., Stamatoyannopoulos J., Nickerson D.A., McCallion A.S., Darnell R., Eichler E.