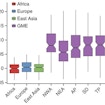
Joseph Gleeson and colleagues have generated a publically accessible whole-exome variome from 1,111 unrelated individuals within the Greater Middle East Variome Consortium.
Highlights of SFARI-funded papers, selected by the SFARI science team.

Joseph Gleeson and colleagues have generated a publically accessible whole-exome variome from 1,111 unrelated individuals within the Greater Middle East Variome Consortium.
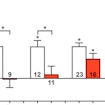
David Ginty shows that four different mice harboring mutations in ASD risk genes have altered peripheral sensory neuron processing that contributes to ASD-like phenotypes.
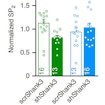
Christian Luscher and Camilla Bellone show that ASD- model mice with reduced SHANK3 in the VTA have altered excitatory synapse transmission and impaired social behaviors.
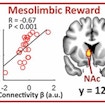
Using fMRI in typically developing children, Vinod Menon finds one’s mother’s voice activates a network of brain regions predictive of a child’s social communication skills.
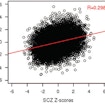
Dan Arking compares large-scale transcriptome datasets from neurotypical individuals and those with autism, schizophrenia and bipolar disorder.
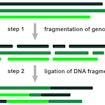
Michael Wigler and Dan Levy develop a new fragmentation and sequencing method for CNV analysis that generates high-resolution CNV data at lower cost than traditional analyses.

Benjamin Philpot shows that seizure susceptibility in an Angelman syndrome mouse model results from the specific loss of UBE3A from GABAergic, but not glutamatergic, neurons.
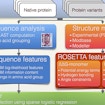
Simons Center for Data Analysis scientist Richard Bonneau develops a method to assess disease gene variants that combines sequence analysis and protein structural modeling.