Benjamin Cheyette finds higher rates of DIXDCI disruptions in individuals with ASD, schizophrenia or bipolar, and DIXDC1 signaling rescue improves mouse ASD-like behaviors.
Highlights of SFARI-funded papers, selected by the SFARI science team.

Benjamin Cheyette finds higher rates of DIXDCI disruptions in individuals with ASD, schizophrenia or bipolar, and DIXDC1 signaling rescue improves mouse ASD-like behaviors.
Rudolf Jaenisch has developed a method enabling the generation of microglia-like cells from human embryonic stem cells and induced pluripotent stem cells.
Liqun Luo uses conditional, cell-type-specific RAI1 deletions in mice to assess how loss of RAI1 contributes to neurodevelopmental phenotypes in Smith-Magenis syndrome.
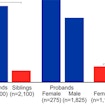
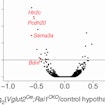
By assessing human accelerated regions (HARs) in healthy individuals and those with ASD, Chris Walsh shows HARs regulate human social and cognitive behavior and ASD risk.
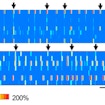

Huda Zoghbi and Stelios Smirnakis investigate how opposing molecular deficits in MeCP2 duplication and Rett syndromes lead to similar behavioral phenotypes.
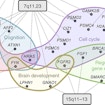
Simons Center for Data Analysis scientist Olga Troyanskaya has developed a machine-learning tool that uses neural interaction networks to predict autism risk genes.
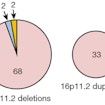
Evan Eichler reconstructs the evolution of chromosome 16p11.2 and finds most disease-causing rearrangements map within a H. sapiens-specific duplication containing BOLA2.
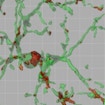
Beth Stevens shows that, in MeCP2 null mice, microglia excessively eliminate presynaptic inputs, targeting synapses previously weakened by MeCP2 loss in other CNS cell types.