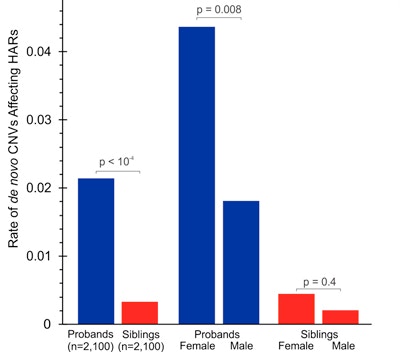
Human accelerated regions (HARs) represent conserved genetic loci that show elevated divergence in humans compared with other species. Such regions have been suggested to play roles in the evolution of human-specific social and cognitive behaviors. In the current study, Christopher Walsh and his colleagues used existing genomic data from healthy individuals to demonstrate essential regulatory functions of HARs, particularly in neural tissues. Next, they investigated whether functional HAR mutations might be linked to deficits in human social or cognitive behaviors. An analysis of copy number variants (CNVs) in the Simons Simplex Collection (SSC) revealed that individuals with autism have an enrichment of rare, de novo HAR-containing CNVs. The researchers then analyzed whole-genome sequencing data from families of Middle Eastern descent with known consanguinity and with one or more children who have autism. The results revealed that rare homozygous mutations in HARs contribute to risk in up to 5 percent of consanguineous autism cases. Functional assessments in neuronal cultures and transgenic mice suggest that many of the identified HAR mutations affect the transcriptional regulation of genes important for neural development. Combined, these data suggest that HARs are essential for normal development and that mutations in HARs significantly contribute to the genetic risk for autism.
Reference(s)
Mutations in human accelerated regions disrupt cognition and social behavior.
Doan R.N., Bae B.I., Cubelos B., Chang C., Hossain A.A., Al-Saad S., Mukaddes N.M., Oner O., Al-Saffar M., Balkhy S., Gascon G.G., Homozygosity Mapping Consortium for Autism, Nieto M., Walsh C.