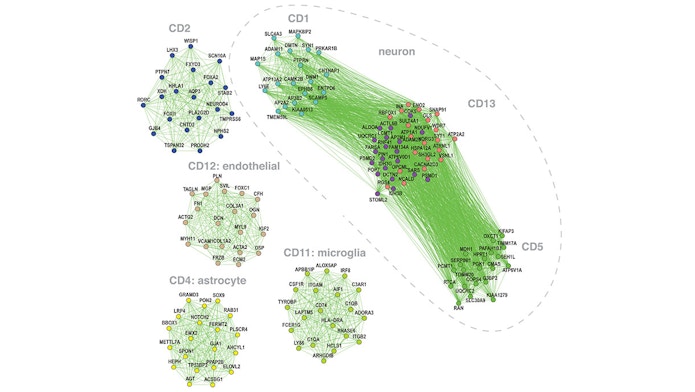
Advances in large-scale genetics efforts have led to the identification of hundreds of genetic risk factors for a variety of psychiatric diseases. Yet how these risk factors function in distinct disorders remains unclear. To address this issue, SFARI Bridge to Independence finalist Michael Gandal, SFARI Investigator Daniel Geschwind and colleagues examined quantitative brain transcriptome profiles for five major psychiatric diseases: autism, schizophrenia, bipolar disorder, depression and alcoholism. Comparison between these profiles uncovered alterations in a variety of shared and distinct gene expression networks. By comparing these profiles to single-nucleotide polymorphism (SNP) risk factors for each disorder, the authors also demonstrated that overlapping alterations in gene-expression profiles mirror the overlap in risk SNPs between disorders. These data argue that these risk SNPs play a causal role in regulating the identified gene pathways and suggest underlying mechanisms by which these risk variants lead to shared behavioral and clinical presentations.
Reference(s)
Shared molecular neuropathology across major psychiatric disorders parallels polygenic overlap.
Gandal M., Haney J.R., Parikshak N.N., Leppa V., Ramaswami G., Hartl C., Schork A.J., Appadurai V., Buil A., Werge T.M., Liu C., White K.P., CommonMind Consortium., PsychENCODE Consortium, iPSYCH-BROAD Working Group., Horvath S., Geschwind D.