The human genome contains hundreds of thousands of enhancers (noncoding regulatory elements that modulate the expression of target genes) yet the discovery and functional characterization of enhancers remains a major challenge.
To address this issue, Xi Chen, Research Scientist at the Flatiron Institute Center for Computational Biology, Olga Troyanskaya, Deputy Director for Genomics at the Flatiron Institute Center for Computational Biology, and their colleagues developed a Bayesian integrative and machine-learning framework called FENRIR (functional enhancer network for relationship inference of regulatory regions). This novel approach integrates thousands of epigenetic and functional genomics data sets to enable the prediction of enhancer function in specific tissues and the prioritization of enhancer-disease associations.
Troyanskaya’s team used FENRIR to build functional networks of 47,881 enhancer regions and 11,447 target genes for 140 human tissues. They then demonstrated the power of the FENRIR networks to prioritize enhancers associated with more than 20 diverse disorders, including cancer and immune and neurological disorders.
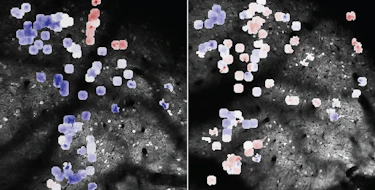
They also performed a case study of autism spectrum disorder (ASD). The FENRIR-predicted ASD-associated enhancers (4,344 in total) were found to be significantly more enriched with proband-specific de novo mutations than sibling-specific mutations based on whole-genome sequencing data from 1,790 Simons Simplex Collection families. Next, Chandra Theesfeld, Research Scientist working with Troyanskaya at Princeton University, selected eight of these candidate ASD-associated enhancers to experimentally validate whether they were able to drive transcriptional activity of target genes in human cells and whether the proband-specific mutations displayed differential activities compared to sibling alleles. All eight of the selected enhancers demonstrated allele-specific regulatory effects, as measured by luciferase reporter assays.
The interactive FENRIR server, with analysis code and pre-generated enhancer networks, is available via this web portal. The researchers plan to update it regularly as new data become available. This bioinformatics tool will be a valuable resource for researchers interested in studying tissue-specific enhancers and their role in disease.
Reference(s)
Tissue-specific enhancer functional networks for associating distal regulatory regions to disease.
Chen X., Zhou J., Zhang R., Wong A.K., Park C.Y., Theesfeld C.L., Troyanskaya O.G.