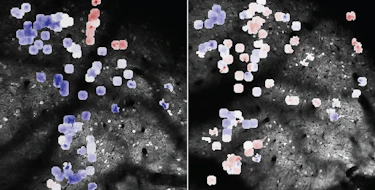
Missense mutations represent one of the greatest interpretive challenges in autism genetics, given the difficulty in determining whether a particular variant is not just functional but also pathogenic. A range of statistical and functional approaches are likely to be needed to inform genetic counselors and clinicians.
Now, SFARI Investigator Paul Sternberg and colleagues have established a pipeline and a suite of assays in C. elegans that can be used to screen and prioritize missense mutations identified in human genetic studies of autism.
In the new work — partly funded by a SFARI targeted award — Sternberg’s team tested 20 autism-associated missense changes in 11 genes (CACNA1D, CHD7, CHD8, CUL3, DLG4, GLRA2, MAPK3, NAA15, PTEN, SYNGAP1 and TPH2). A range of quantitative assays were developed to test for functional defects imparted by these variants, including body morphology (length, width and area), movement (speed, reversal rate and sinusoidal wavelength and amplitude) and fecundity. While all 20 of the missense variants were predicted to perturb protein function, only 14 showed a detectable phenotype in one of the functional assays, suggesting that they should receive priority for follow-up.
Undoubtedly, a larger range of assays in multiple species will have the greatest predictive power, and additional studies by SFARI Investigators Kurt Haas (McDiarmid et al., bioRxiv, 2019) and Haiyuan Yu (Chen et al., Nat. Genet., 2018) are contributing to this effort.

Reference(s)
Autism-associated missense genetic variants impact locomotion and neurodevelopment in Caenorhabditis elegans.
Wong W.R., Brugman K.I., Maher S., Oh J.Y., Howe K., Kato M., Sternberg P.