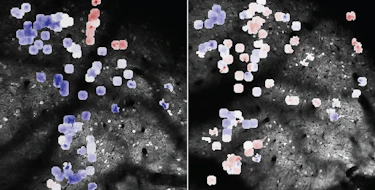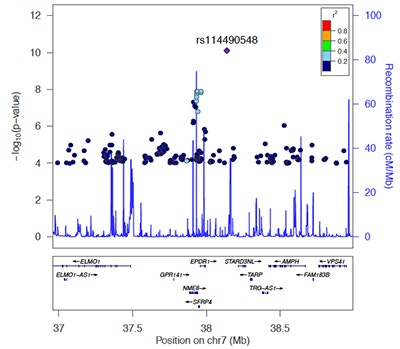
Interactions between genes, referred to as epistasis, have been shown to play a large role in the variation in complex traits in model organisms. Yet human genome-wide association studies (GWAS) have had limited success in identifying genetic interactions. SFARI Investigator Lauren Weiss and colleagues investigated whether analyses of a known autism spectrum disorder (ASD)-associated signaling pathway, the Ras/MAPK pathway, might provide insights into genetic epistasis in idiopathic ASD. Weiss’s team generated an ASD GWAS dataset from available cohorts, including the Simons Simplex Collection, and found that common single nucleotide polymorphisms (SNPs) in RAS/MAPK genes are significantly enriched for association with ASD. They then performed a genome-wide screen for interactors with Ras/MAPK gene SNPs and found 19 unique epistatic pairs that met criteria for significance in ASD cases compared with controls. They also performed a modifier screen based on quantifiable measures of a social responsiveness trait in individuals with rare RASopathies – Mendelian disorders of the Ras/MAPK pathway. They went on to show that the expression of GPR141, one of the genes located in one of the identified epistatic loci, was reduced in RASopathy neural cell lines. This study highlights that a reverse pathway genetic approach, focusing on key biological pathways, can be applied to study epistasis in ASD.
Reference(s)
Reverse pathway genetic approach identifies epistasis in autism spectrum disorders.
Mitra I., Lavillaureix A., Yeh E., Traglia M., Tsang K., Bearden C., Rauen K.A., Weiss L.